Sabina Sara Pfister
Neuroscientists, Software Developer, Open Source Enthusiast
DYNNETWORK: Interactive Dynamic Networks
Description
Analysis of dynamical information (gene expression, protein interactions, etc...) is become of increasing interest for the scientific community, and the development of open source tools to provide the ability of analyzing such data sets is highly required. Cytoscape offers an ideal and already well developed framework into which network structured data can be accessed, visualized, and analyzed.
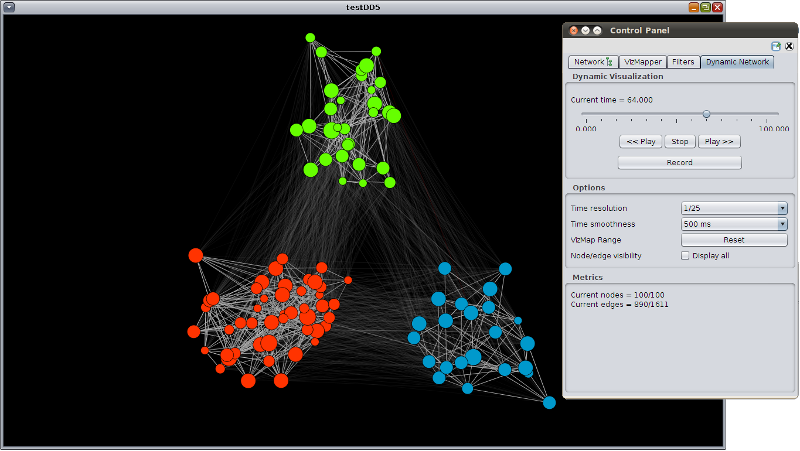
DynNetwork is a OSGi bundle to provide support for importing, visualizing, and analysing dynamical networks in Cytoscape 3.0. Dynamical networks are imported in form of XGMML files, and the dynamics of nodes, edges, attributes can be visualized at the desired time points. To improve visualization, the nodes can be animated with a dynamic force layout algorithm.
Source code
Code and instruction are available here.